Biomedical researchers plan to create the Pediatric Cell Atlas (PCA), a powerful new resource for fine-grained scientific understanding of human growth and development. Drawing on dramatic recent advances in technology, the Atlas will offer an unprecedented window into the unique biology of children by benchmarking healthy and abnormal tissues at the level of single cells — the basic units of biology.
“Pediatricians are familiar with the mantra that ‘children are not just small adults,’” said first author Deanne M. Taylor, PhD, Director of Bioinformatics in the Department of Biomedical and Health Informatics at Children’s Hospital of Philadelphia (CHOP) and a research assistant professor of Pediatrics in the Perelman School of Medicine at the University of Pennsylvania. “Children’s diseases, symptoms, outcomes and therapies are often age-dependent, along with differences in physiology, presentation and drug responses compared to those occurring in adults. With this Atlas, we’ll have a standard reference tool showing, at different ages, which cells are doing the work for a child to grow healthily.”
Taylor is the first and corresponding author of an open-access perspective article on the PCA in the journal Developmental Cell, online March 28. Her co-first authors are Bruce J. Aronow, PhD, of the Division of Biomedical Informatics at Cincinnati Children’s Hospital Medical Center, and CHOP researcher Kai Tan, PhD, also of the University of Pennsylvania. Other co-authors represent Boston Children’s Hospital, Children’s National Medical Center, Beatrix Children’s Hospital and the National Institutes of Health, among several other institutions. The PCA, added Taylor, aims to bring together a global coalition of pediatric researchers to create this high-definition view of children’s health.
Most parents are familiar with growth charts, on which a pediatrician periodically logs a child’s height and weight in comparison to national averages. The PCA will compile age-matched trajectories of tissue and organ development in healthy children, referencing key data patterns in cell differentiation and cell signaling. Those trajectories will offer a standard for researchers to better understand when and how childhood illnesses diverge from those patterns, because of genetic influences, environmental factors or both.
Overall, support for pediatric research lags behind support for adult health research, with a corresponding lag in breakthrough biomedical discoveries and translation into clinical treatments for children.
While addressing this disparity with its own robust pediatric research program, CHOP also places its work in a broader context of lifespan medicine — recognizing the progression of health and disease over a fetal-to-adult continuum. Many lifelong chronic diseases, such as diabetes, asthma and neuropsychiatric conditions, first become apparent during childhood or adolescence, so designing or improving interventions during childhood could yield lifelong benefits. The PCA would provide key evidence for advancing such efforts.
Historically, researchers have extracted much cellular and genetic data from bulk tissue samples through measuring levels of active genes and proteins. The available bulk data measurements are low-resolution in their ability to capture the diversity of cell types, cell tasks and cell stages. “We can’t assume that all the cells in one tissue, or even one section of a tissue, are doing the same jobs,” said Taylor. “Some cell types, such as stem cells, may be present in very low numbers, but may be performing key tasks. Cells may also perform different tasks at different stages, but these details may be lost in the noise from bulk data.”
Single-cell analyses, in contrast, separate tissues into individual cells, and analyze each cell’s molecular signature. Technologies that have become available over the past few years combine next-generation sequencing and massively parallel processing, such as RNA sequencing, of single cells. Single-cell studies, said Taylor, open up a high-definition view of cell physiology and functioning that will expand scientific knowledge of health and disease — particularly during the dynamic childhood period, when growth and development are under way.
The PCA will be part of a broader international consortium, the Human Cell Atlas (HCA), which held its initial planning meeting in 2016. Two of the co-founders of the HCA initiative, Dr. Aviv Regev of the Broad Institute, and Dr. Sarah Teichmann, of the Wellcome Sanger Institute, are co-authors of the current PCA paper.
The PCA will share data among members and with other researchers worldwide. It will also store data in the HCA’s repository, said Taylor, with associated biobanking and data repositories in different centers available to biological researchers. As the PCA moves forward, it will develop its overall organization, data systems and multiple projects, including pilot studies of specific organs and disease areas, such as childhood cancers. It will also compare atlas data from healthy tissues with data from diseased tissue, such as those generated by the Pediatric Tumor Atlas, sponsored by the Cancer Moonshot of the National Cancer Institute.
“Ultimately, researchers would leverage knowledge from single-cell data into a deeper understanding of organ development and function, to better inform precision treatments to advance children’s health,” added Taylor.
Deanne M. Taylor et al, “The Pediatric Cell Atlas: Defining the growth phase of human development at single-cell resolution,” Developmental Cell, online March 28, 2019.
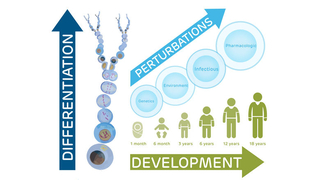
Photo: The PCA will detail how cells and tissues change during childhood development, especially when influenced by genes, infections and the environment.
Featured in this article
Specialties & Programs
Biomedical researchers plan to create the Pediatric Cell Atlas (PCA), a powerful new resource for fine-grained scientific understanding of human growth and development. Drawing on dramatic recent advances in technology, the Atlas will offer an unprecedented window into the unique biology of children by benchmarking healthy and abnormal tissues at the level of single cells — the basic units of biology.
“Pediatricians are familiar with the mantra that ‘children are not just small adults,’” said first author Deanne M. Taylor, PhD, Director of Bioinformatics in the Department of Biomedical and Health Informatics at Children’s Hospital of Philadelphia (CHOP) and a research assistant professor of Pediatrics in the Perelman School of Medicine at the University of Pennsylvania. “Children’s diseases, symptoms, outcomes and therapies are often age-dependent, along with differences in physiology, presentation and drug responses compared to those occurring in adults. With this Atlas, we’ll have a standard reference tool showing, at different ages, which cells are doing the work for a child to grow healthily.”
Taylor is the first and corresponding author of an open-access perspective article on the PCA in the journal Developmental Cell, online March 28. Her co-first authors are Bruce J. Aronow, PhD, of the Division of Biomedical Informatics at Cincinnati Children’s Hospital Medical Center, and CHOP researcher Kai Tan, PhD, also of the University of Pennsylvania. Other co-authors represent Boston Children’s Hospital, Children’s National Medical Center, Beatrix Children’s Hospital and the National Institutes of Health, among several other institutions. The PCA, added Taylor, aims to bring together a global coalition of pediatric researchers to create this high-definition view of children’s health.
Most parents are familiar with growth charts, on which a pediatrician periodically logs a child’s height and weight in comparison to national averages. The PCA will compile age-matched trajectories of tissue and organ development in healthy children, referencing key data patterns in cell differentiation and cell signaling. Those trajectories will offer a standard for researchers to better understand when and how childhood illnesses diverge from those patterns, because of genetic influences, environmental factors or both.
Overall, support for pediatric research lags behind support for adult health research, with a corresponding lag in breakthrough biomedical discoveries and translation into clinical treatments for children.
While addressing this disparity with its own robust pediatric research program, CHOP also places its work in a broader context of lifespan medicine — recognizing the progression of health and disease over a fetal-to-adult continuum. Many lifelong chronic diseases, such as diabetes, asthma and neuropsychiatric conditions, first become apparent during childhood or adolescence, so designing or improving interventions during childhood could yield lifelong benefits. The PCA would provide key evidence for advancing such efforts.
Historically, researchers have extracted much cellular and genetic data from bulk tissue samples through measuring levels of active genes and proteins. The available bulk data measurements are low-resolution in their ability to capture the diversity of cell types, cell tasks and cell stages. “We can’t assume that all the cells in one tissue, or even one section of a tissue, are doing the same jobs,” said Taylor. “Some cell types, such as stem cells, may be present in very low numbers, but may be performing key tasks. Cells may also perform different tasks at different stages, but these details may be lost in the noise from bulk data.”
Single-cell analyses, in contrast, separate tissues into individual cells, and analyze each cell’s molecular signature. Technologies that have become available over the past few years combine next-generation sequencing and massively parallel processing, such as RNA sequencing, of single cells. Single-cell studies, said Taylor, open up a high-definition view of cell physiology and functioning that will expand scientific knowledge of health and disease — particularly during the dynamic childhood period, when growth and development are under way.
The PCA will be part of a broader international consortium, the Human Cell Atlas (HCA), which held its initial planning meeting in 2016. Two of the co-founders of the HCA initiative, Dr. Aviv Regev of the Broad Institute, and Dr. Sarah Teichmann, of the Wellcome Sanger Institute, are co-authors of the current PCA paper.
The PCA will share data among members and with other researchers worldwide. It will also store data in the HCA’s repository, said Taylor, with associated biobanking and data repositories in different centers available to biological researchers. As the PCA moves forward, it will develop its overall organization, data systems and multiple projects, including pilot studies of specific organs and disease areas, such as childhood cancers. It will also compare atlas data from healthy tissues with data from diseased tissue, such as those generated by the Pediatric Tumor Atlas, sponsored by the Cancer Moonshot of the National Cancer Institute.
“Ultimately, researchers would leverage knowledge from single-cell data into a deeper understanding of organ development and function, to better inform precision treatments to advance children’s health,” added Taylor.
Deanne M. Taylor et al, “The Pediatric Cell Atlas: Defining the growth phase of human development at single-cell resolution,” Developmental Cell, online March 28, 2019.

Photo: The PCA will detail how cells and tissues change during childhood development, especially when influenced by genes, infections and the environment.
Contact us
Department of Biomedical and Health Informatics (DBHi)