Brain Imaging Links Language Delay to Chromosome Deletion in Children with Neurological Disorders
CHOP-Led MEG Study Suggests Pathway Common to Autism, Other Disorders; Hints at Drugs to Target Overlapping Illnesses
Published on
Published on
Children born with a DNA abnormality on chromosome 16 already linked to neurodevelopmental problems show measurable delays in processing sound and language, says a study team of radiologists and psychologists.
By strengthening the case that the deleted gene disrupts a key biological pathway, the research may lay the foundation for future medical treatments for specific subtypes of autism, along with cognitive and language disabilities.
“This study shows an important connection between gene differences and differences in neurophysiology,” said study leader Timothy P.L. Roberts, PhD, vice chair of Radiology Research at The Children’s Hospital of Philadelphia (CHOP) and a researcher at CHOP’s Center for Autism Research. “It may also help to bridge a largely unexplored gap between genetics and behavior.”
Roberts, who holds the Oberkircher Family Endowed Chair in Pediatric Radiology at CHOP, led the study published online Feb. 11 in Cerebral Cortex, collaborating with a group led by Elliott H. Sherr, MD, PhD, of the University of California, San Francisco (UCSF).
 The researchers examined children with copy number variants—either deletions or duplications of DNA— at the genetic site 16p11.2. Previous researchers had found that this location on chromosome 16 was associated with a subset of autism spectrum disorders (ASDs) and with language impairments and developmental delays.
The researchers examined children with copy number variants—either deletions or duplications of DNA— at the genetic site 16p11.2. Previous researchers had found that this location on chromosome 16 was associated with a subset of autism spectrum disorders (ASDs) and with language impairments and developmental delays.
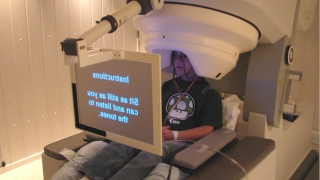
Roberts and colleagues used magnetoencephalography (MEG), which detects magnetic fields in the brain, just as electroencephalography (EEG) detects electrical fields. As each child heard a series of tones, the MEG machine analyzed changing magnetic fields in the child’s brain, measuring an auditory processing delay called the M100 response latency.
The researchers analyzed 115 children: 43 with the 16p11.2 deletion, 23 with the 16p11.2 duplication, and 49 healthy controls. The children were from two centers, CHOP and UCSF. Only a fraction of the children had ASD diagnoses: 11 of the 43 with the deletion, and 2 of the 23 with the duplication.
In children with the deletion, the researchers found a significant delay: 23 milliseconds (ms), a figure that Roberts called “stunningly high” compared to the healthy children. There was no such delay among children with the duplication, who actually had a non-significant tendency to process sounds faster than the control subjects.
The 23-ms delay, about one-fortieth of a second, was twice as high as the 11-ms M100 delay that Roberts found in a 2010 MEG study of children with ASDs. In that study, Roberts remarked that 11 milliseconds is a brief interval, but that it meant that a child hearing the word ‘elephant’ would still be processing the ‘el’ sound while other children moved on, with delays cascading as a conversation progresses.
While the 2010 study focused on children diagnosed with ASDs, Roberts added, the current study took a “genetics first” approach, analyzing children known to have genetic variants with or without ASD diagnoses. “We have approached the problem from both ends,” he said. The previous study found a link between the brain and behavior, while this new study found a link between genetics and the brain.”
Although not all of the children with CNVs had autism, all of them had some neurological or learning disabilities, he added. Because the severity of neurodevelopmental symptoms did not correlate with the length of the auditory processing delay, the M100 delay may not become a clear-cut diagnostic biomarker in neurological disorders, but it may be a clue to an important common pathway in neurobiology.
“We don’t yet know the significance of the 23-millisecond delay, but we have established its origin in genetics,” Roberts said. “It seems to be a proxy for something of biological significance.” He noted that this finding meshes with a current research strategy articulated by the National Institutes of Health—focusing on research domains criteria in neurological diseases. Because ASDs and mental disorders are very diverse and heterogeneous, involving many different genes, a research domains approach seeks to discover genes that may overlap differing disorders but operate on common biological pathways and processes.
Further studies, said Roberts, will investigate other genes previously implicated in ASDs and other psychiatric disorders, to determine whether they also involve M100 response delays. “Our goal is to unify diverse genes along a few common pathways, some of which may be treatable with specific therapies,” said Roberts.
He added that his laboratory is planning a very small pilot study of children with ASDs who have the M100 response latency. Using a drug that acts on synaptic transmissions (signals across nerve cells), he will analyze whether this drug reduces the M100 auditory delays.
This research is at an early stage, Roberts stressed, adding that the biological mechanisms underlying the chromosome deletion and auditory delays remain undiscovered. “We don’t know, for instance, whether the abnormality that leads to the delay happens at the synapses or in the brain’s white matter, which acts as highway for carrying brain signals. Our next studies may help answer that question.”
In addition to his CHOP title, Roberts also is on the faculty of the Perelman School of Medicine at the University of Pennsylvania.
The Simons Foundation provided grant support to Roberts for this study. John E. Spiro of the Simons Foundation was a co-author of the study.
“Auditory Evoked M100 Response Latency Is Delayed in Children with 16p11.2 Deletion but not 16p11.2 Duplication,” Cerebral Cortex, published Feb. 11, 2015.
Post Categories: CHOP News, Chromosome Deletions